Overview & Concept
What problem does OncoMetabolismGPS solve?
Cancer metabolic reprogramming is a highly complex phenomenon driven by multiple molecular dimensions. Traditionally, omic layers such as copy number variations, DNA methylation, mRNA expression, miRNA expression, protein abundance, and mutations are often analyzed separately. This fragmented view can obscure the broader metabolic regulatory circuitries that shape tumor progression, immune contexture, and clinical behavior.
OncoMetabolismGPS addresses this challenge by providing an integrated pan-cancer atlas of metabolic regulatory circuitries. Reflecting the conceptual framework of the preprint A Pan-Cancer Atlas of Metabolic Regulatory Circuitries Integrating Multi-Omic, Immune, and Clinical Dimensions, the platform enables users to explore multi-omic signatures together with regulatory interactions, immune features, and clinical associations within a single interactive environment.
How to navigate the platform
Use the navigation bar above to explore different dimensions of the OncoMetabolismGPS resource:
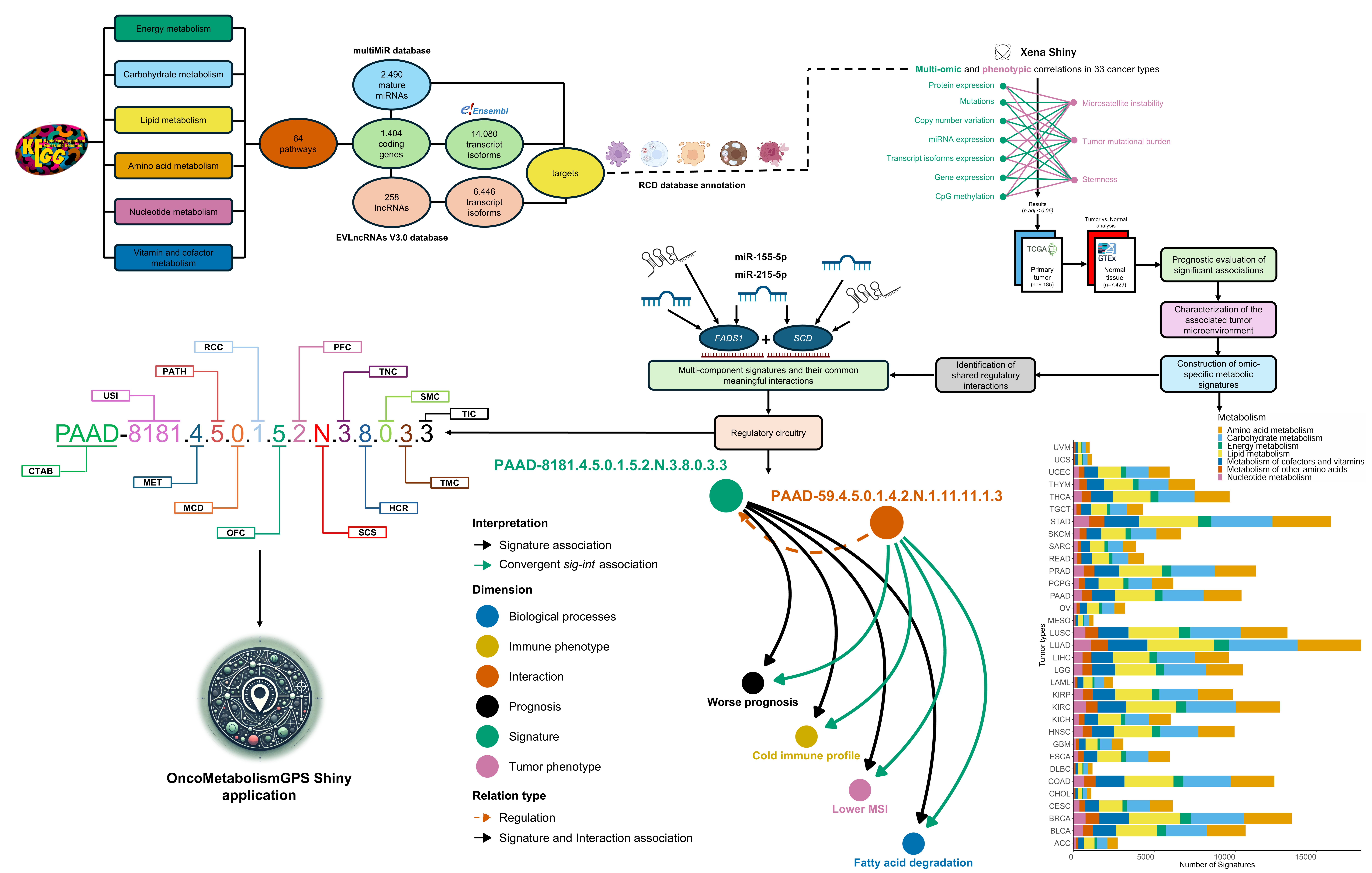
- Framework View the graphical summary of the computational pipeline used to generate the atlas and access the interactive components of the platform.
- Dashboard Search for a gene, miRNA, or lncRNA to identify associated signatures, inspect biological interpretation, and explore regulatory relationships linked to your target.
- Browse Signatures Explore signatures across omic layers, including CNV, mRNA, methylation, mutation, miRNA, protein, and transcript-based signatures.
- Regulatory Circuitries Visualize meaningful metabolic regulatory circuitries and inspect how different molecular entities connect with biological and clinical dimensions.
- User Manual Access guidance on how to interpret the platform outputs and navigate the available analytical resources.
- Citation Find the recommended citation for the OncoMetabolismGPS Shiny App and the associated preprint.
Conceptual foundation of the resource
OncoMetabolismGPS was designed to move beyond single-layer analyses by integrating metabolic pathway context, multi-omic signatures, regulatory interactions, immune dimensions, and clinical outcomes. This structure allows users to distinguish convergent and divergent metabolic circuitries and to prioritize candidate biomarkers and therapeutic targets from a systems-level perspective.
You will obtain: matching nomenclatures, a biological interpretation of the selected entry, and its associated regulatory network.
Start with a gene, miRNA or lncRNA to retrieve related entries available in the platform.
Browse the matched nomenclatures and choose the entry you want to inspect in more detail.
Read the interpretation of the selected result and inspect its regulatory circuitry in the network panel.
Download Data Download Interpretation
Download Plot (PDF)
Integrative Summary
Table results
Integrative Summary
Table results
Integrative Summary
Table results
Integrative Summary
Table results
Integrative Summary
Table results
Integrative Summary
Table results
Integrative summary for the selected interaction
Regulatory circuitry network
All interactions for this Nomenclature
Click a row to update the textual summary above.
OncoMetabolismGPS is an interactive platform for exploring multi-omic metabolic signatures, their regulatory relationships, immune context, and clinical associations across 33 cancer types.
- Search a gene, miRNA, or lncRNA and retrieve associated signatures.
- Inspect structured biological interpretation for each nomenclature.
- Visualize regulatory circuitries when available.
- Browse signatures by omic layer and biological filters.
- Access audio and video content for a guided introduction to the resource.
Use the Dashboard when you want to start from a specific molecular entity and discover how it appears across signatures and regulatory contexts.
Step 1 Enter a target – type a gene symbol, mature miRNA, or lncRNA in the search field.
Step 2 Run the query – click the search button.
A status message indicates whether the target is present in the database.
Nomenclatures – all signatures containing the queried target.
Interpretation – structured summary of the selected nomenclature.
Regulatory circuitry – network visualization when regulatory information is available.
- Click a nomenclature entry in the results panel.
- The interpretation panel updates with tumor type, omic and phenotype layers, metabolism, pathway, regulated cell death, prognosis, and immune profile.
- If a regulatory network exists, the circuitry panel displays the corresponding graph.
-
Download Data
exports the selected row as a
.csvfile. -
Download Interpretation
exports a text summary as a
.txtfile. - Download Plot (PDF) exports the circuitry plot when available.
Do I need a nomenclature code before searching?
No. You begin with the biological target, and the app retrieves the related nomenclatures automatically.
Can I inspect more than one nomenclature?
Yes. Each time you click a different nomenclature, the interpretation and circuitry panels update accordingly.
Why do some signatures show no circuitry plot?
Some nomenclatures do not yet have a consolidated regulatory circuitry available in the current version of the resource.
Use this section when you want to explore the atlas systematically rather than starting from one target.
- Omic layer – defined by the active tab.
- Cancer type – TCGA tumor abbreviation.
- Metabolism – broad metabolic class.
- Pathway – specific metabolic route.
- Regulated cell death – biological death program linked to the signature.
- Signature – the final matching nomenclature.
- Integrative summary of the selected signature.
- Tumor and non-tumor association metrics.
- Survival-related information when available.
- Immune and microenvironment-related annotations.
- Exportable tables for downstream analysis.
This section helps you inspect how molecular entities are connected within meaningful metabolic regulatory circuitries.
- Use it to explore interaction structure beyond isolated signatures.
- Inspect how regulators, metabolic processes, and clinical or biological dimensions are linked.
- Use this area for systems-level interpretation of the atlas.
The multimedia area offers a guided introduction to the project for users who prefer audio or video explanations before interacting with the data.
- Podcast – concise explanation of the rationale and key ideas behind the platform.
- Presentation video – walkthrough of the resource and selected use cases.
- Start with Framework to see the global computational logic.
- Move to Overview & Concept to understand the biological rationale.
- Use Dashboard when you already have a target of interest.
- Use Browse Signatures for systematic exploratory analysis.
- Consult Regulatory Circuitries for deeper systems-level interpretation.
- Use Multimedia for a guided introduction.
Discover the Spatial Architecture of Metabolism
While OncoMetabolismGPS provides a comprehensive view of metabolic regulators across clinical and immune dimensions, the structural relationships driving these networks exist in complex, multidimensional spaces. We invite you to explore our companion platform, SigPolytope.
SigPolytope projects dense metabolic regulatory circuitries into an interactive, pan-cancer geometric atlas. By transforming multi-omic topologies into multidimensional spaces, you can visually explore structural similarities, identify distinct spatial architectures, and navigate tumor metabolism in a completely reimagined way.
Launch the SigPolytope Geometric AtlasMeet the research team behind OncoMetabolismGPS.

Logo concept by Higor Almeida Cordeiro Nogueira using ChatGPT.
For questions, suggestions or technical issues, contact higoralmeida1995@gmail.com .
How to Cite
If you use the OncoMetabolismGPS Shiny App or the results generated by this application, please cite the published article:
Almeida Cordeiro Nogueira, H.; Rodrigues de Souza, E.; dos Santos Lopes, V.; Medina-Acosta, E. (2026). A pan-cancer atlas of metabolic regulatory circuitries integrating multi-omic, immune, and clinical dimensions. Frontiers in Molecular Biosciences , 13, 1845099. DOI: 10.3389/fmolb.2026.1845099
BibTeX
@article{Nogueira2026OncoMetabolismGPS,
title={A pan-cancer atlas of metabolic regulatory circuitries integrating multi-omic, immune, and clinical dimensions},
author={Almeida Cordeiro Nogueira, Higor and Rodrigues de Souza, Emanuell and dos Santos Lopes, Victor and Medina-Acosta, Enrique},
year={2026},
journal={Frontiers in Molecular Biosciences},
volume={13},
pages={1845099},
doi={10.3389/fmolb.2026.1845099},
url={https://doi.org/10.3389/fmolb.2026.1845099}
}